Help for MViewEMA server
Input of MViewEMA server
-
(Mandatory) At least 1 protein complex structure model in PDB format
Users can click the button "Add model" to add text boxes for input more structure models of same protein complex The server page is set to receive up to 10 structure models. - If you choose to upload a zip file containing all models of the same protein (each model file ends with ".pdb"), please compress the folder to ./files or ./dir/files.
- (Optional) Provide your email address
- (Optional) Assign your job name
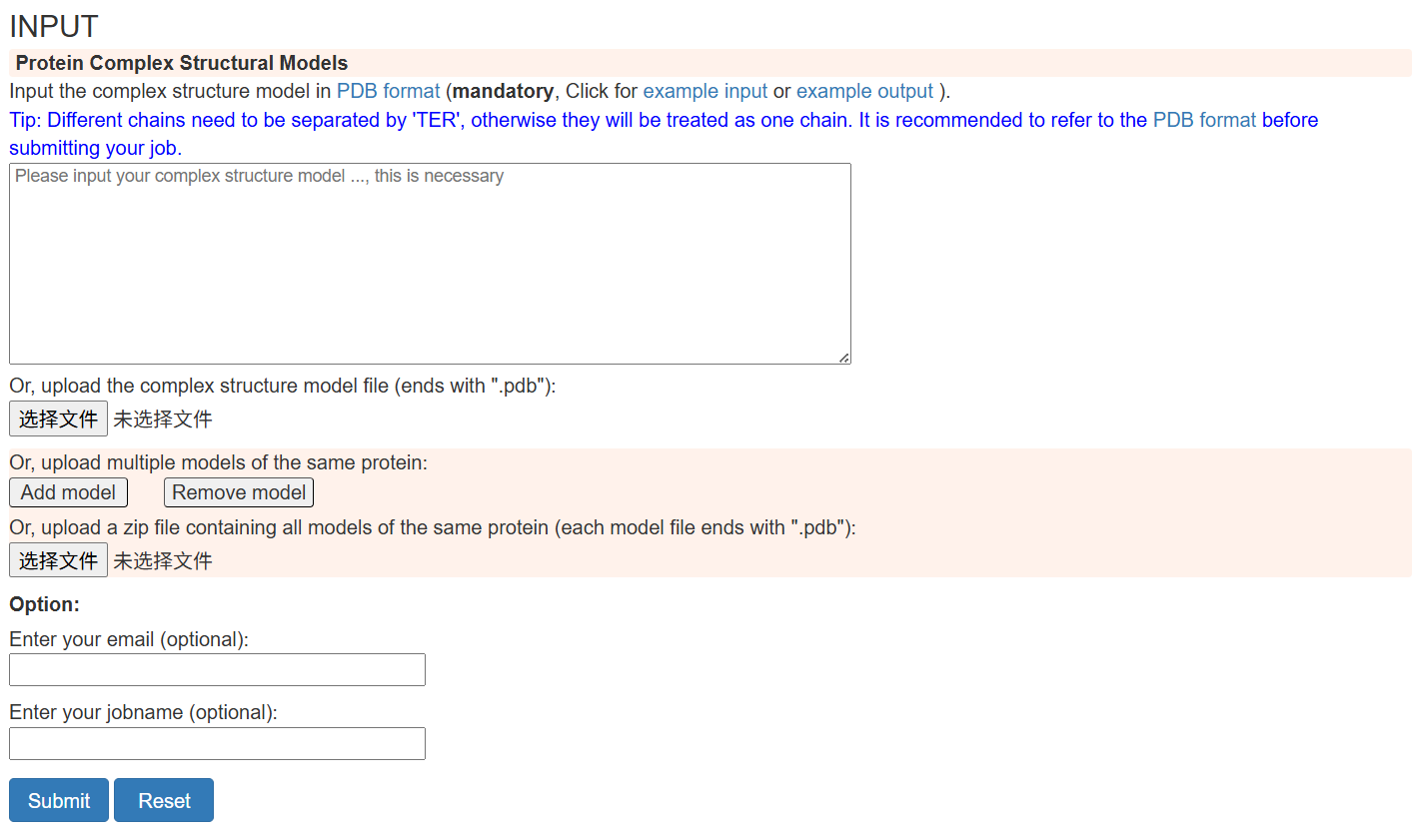 Figure 1. The "Submit" section of complex structure assessment in MViewEMA home page.
Figure 1. The "Submit" section of complex structure assessment in MViewEMA home page.
Output of MViewEMA server
Complex structure assessment:
- The results in the CSV file include the global confidence score predicted for each model.
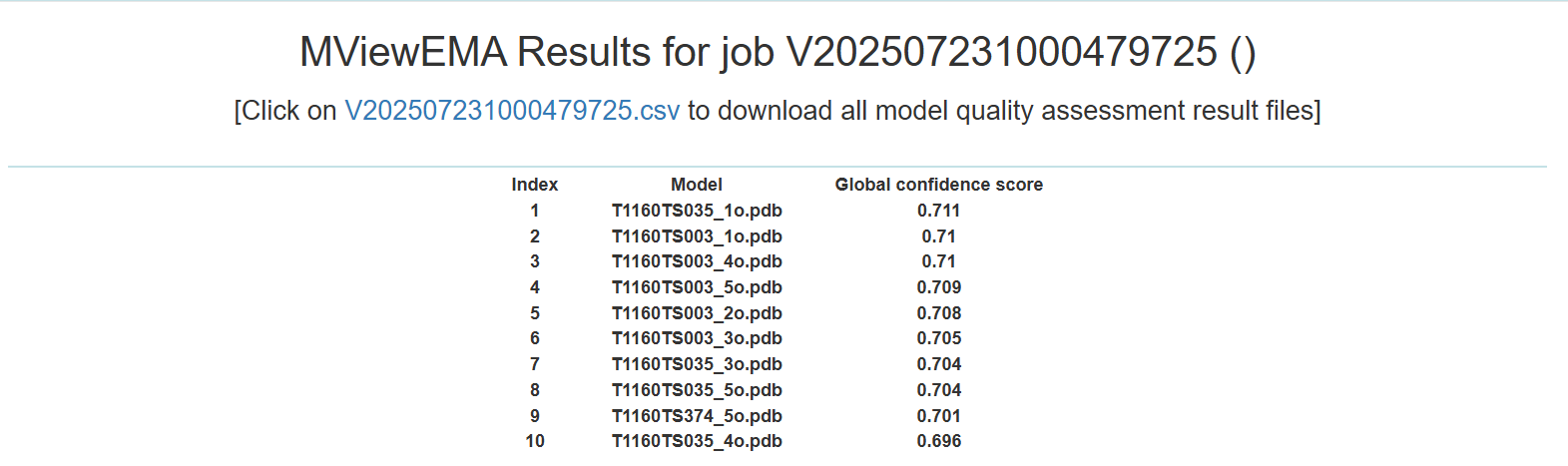 Figure 2. The output of complex structure assessment in MViewEMA result page.
Figure 2. The output of complex structure assessment in MViewEMA result page.
Evaluation Metrics
MViewEMA assesses the structural quality of protein models using TM-score, a widely adopted metric that reflects overall fold accuracy.
- TM-score (Zhang and Skolnick 2004) were calculated using US-align (Zhang et al. 2022) to assess protein single-chain and complex topological similarity.
How to cite MViewEMA?
Please cite the following articles when you use the MViewEMA server:
- Dong Liu, Jun Liu, Haodong Wang, Fang Liang, Guijun Zhang*. DeepUMQA-X: Comprehensive and insightful estimation of model accuracy for protein single-chain and complex. Nucleic Acids Research, 2025, W1(7), W219–W227.
- Dong Liu✝, Biao Zhang✝, Jun Liu, Hui Li*, Le Song* and Guijun Zhang*. GraphCPLMQA: Assessing protein model quality based on deep graph coupled networks using protein language model. Briefings in Bioinformatics, 2023, 25(1):bbad420.
- Saisai Guo✝, Jun Liu✝, Xiaogen Zhou, Guijun Zhang*. DeepUMQA: Ultrafast Shape Recognition-based Protein Model Quality Assessment using Deep Learning. Bioinformatics, 2022, 38(7): 1895-1903.
Need more help?
If you have more questions or comments about the server, please email guijunlab06@163.com.